myBaits Expert –Plant–Angiosperms353
低起始量 - 1 ng-3 μg,推荐100-500 ng
覆盖广 - 捕获覆盖整个目标序列
兼容性 - 兼容NGS文库制备,允许多个碱基错配
- 产品简介
- 工作流程
- 实验数据
- 引用文献
myBaits Angiosperms 353是从600种被子植物中,筛选353个单拷贝核基因,作为捕获序列,利用k-medoid聚类法设计靶向捕获探针,生物素化的RNA探针长度120 bp,对被子植物的353个单拷贝核基因进行定向捕获测序。平均每个物种覆盖283个基因,所有被子植物至少覆盖100个直系同源单拷贝核基因。每个物种至少靶向137 kbp,最大捕获区域为250 kbp,另外覆盖了212 kbp的非编码区。
产品特点:
低起始量 - 1 ng-3 μg,推荐100-500 ng
覆盖广 - 捕获覆盖整个目标序列
兼容性 - 兼容NGS文库制备,允许多个碱基错配
高精度 - 近缘和远缘的个体或者种群捕获,不引入biases
低投入 - 节约时间成本和经费成本
高通量 - 多个文库同一反应进行,每个反应pooling 2-16个样本
通用性 - 适用于被子植物的系统进化分析,建立DNA条形码
产品应用:
系统发生学
进化关系的分析
内含子测序
发现可变区域
DNA Barcoding
对目标序列的富集程度进行评价
产品列表:
| 产品名称 | 规格 | 货号 |
| myBaits Expert Angiosperms 353 v1 | 8 Reactions | 308108.v5 |
| 48 Reactions | 308148.v5 | |
| 96 Reactions | 308196.v5 |
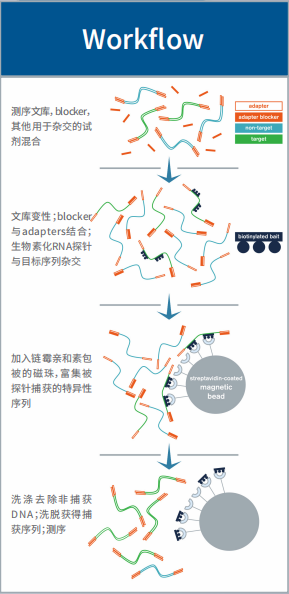
解决方案
我们提供的不仅仅是Angiosperms 353产品,而是基于靶向捕获测序技术的基因组学整体解决方案!
完整流程:
DNA提取→纯度和浓度检测→DNA片段化→测序文库制备→文库质检→杂交捕获→捕获序列富集→PCR产物纯化→PCR产物质检→测序→生物信息学分析。
基因覆盖效率
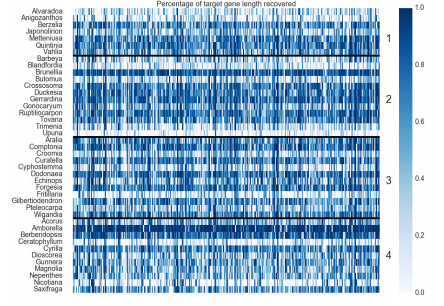
Figure 1. Heatmap of Gene Recovery Efficiency. Each row is one sample, and each column is one gene. Colors indicate the percentage of the target length (calculated by the mean length of all k-medoid transcripts for each gene) recovered. Numbers indicate the Input Category (see main text).
Antonelli, A., Clarkson, J.J., Kainulainen, K., Maurin, O., Brewer, G.E., Davis, A.P., … Baker, W.J. (2021). Settling a family feud: a high-level phylogenomic framework for the Gentianales based on 353 nuclear genes and partial plastomes. American Journal of Botany 108, 1143–1165.
Baker, W.J., Bailey, P., Barber, V., Barker, A., Bellot, S., Bishop, D., … Forest, F. (2021). A Comprehensive Phylogenomic Platform for Exploring the Angiosperm Tree of Life. Systematic Biology.
Brewer, G.E., Clarkson, J.J., Maurin, O., Zuntini, A.R., Barber, V., Bellot, S., … Baker, W.J. (2019). Factors Affecting Targeted Sequencing of 353 Nuclear Genes From Herbarium Specimens Spanning the Diversity of Angiosperms. Front. Plant Sci. 0.
Buerki, S., Callmander, M.W., Acevedo-Rodriguez, P., Lowry, P.P., Munzinger, J., Bailey, P., … Forest, F. (2021). An updated infra-familial classification of Sapindaceae based on targeted enrichment data. American Journal of Botany 108, 1234–1251.
Clarkson, J.J., Zuntini, A.R., Maurin, O., Downie, S.R., Plunkett, G.M., Nicolas, A.N., … Baker, W.J., (2021). A higher-level nuclear phylogenomic study of the carrot family (Apiaceae). American Journal of Botany 108, 1252–1269.
Gaynor, M.L., Fu, C.-N., Gao, L.-M., Lu, L.-M., Soltis, D.E., Soltis, P.S. (2020). Biogeography and ecological niche evolution in Diapensiaceae inferred from phylogenetic analysis. Journal of Systematics and Evolution 58, 646–662.
Hendriks, K.P., Mandáková, T., Hay, N.M., Ly, E., Huysduynen, A.H. van, Tamrakar, R., … Bailey, C.D. (2021). The best of both worlds: Combining lineage-specific and universal bait sets in target-enrichment hybridization reactions. Applications in Plant Sciences 9.
Johnson, M., Pokorny, L., Dodsworth, S., Botigue, L.R., Cowan, R.S., Devault, A., … Wickett, N. (2019). A Universal Probe Set for Targeted Sequencing of 353 Nuclear Genes from Any Flowering Plant Designed Using k-medoids Clustering. Systematic Biology 68(4): 594-606.
Larridon, I., Villaverde, T., Zuntini, A.R., Pokorny, L., Brewer, G.E., Epitawalage, N., … Baker, W.J. (2020). Tackling Rapid Radiations With Targeted Sequencing. Frontiers in Plant Science 10: 1.
Larridon, I., Zuntini, A.R., Barrett, R.L., Wilson, K.L., Bruhl, J.J., Goetghebeur, P., … Roalson, E.H. (2021). Resolving generic limits in Cyperaceae tribe Abildgaardieae using targeted sequencing. Botanical Journal of the Linnean Society 196, 163–187.
Larridon, I., Zuntini, A.R., Léveillé-Bourret, É., Barrett, R.L., Starr, J.R., Muasya, A.M., … Baker, W.J. (2021). A new classification of Cyperaceae (Poales) supported by phylogenomic data. Journal of Systematics and Evolution 59, 852–895.
Lee, A.K., Gilman, I.S., Srivastav, M., Lerner, A.D., Donoghue, M.J., Clement, W.L. (2021). Reconstructing Dipsacales phylogeny using Angiosperms353: issues and insights. American Journal of Botany 108, 1122–1142.
Maurin, O., Anest, A., Bellot, S., Biffin, E., Brewer, G., Charles-Dominique, T., … Lucas, E. (2021). A nuclear phylogenomic study of the angiosperm order Myrtales, exploring the potential and limitations of the universal Angiosperms353 probe set. American Journal of Botany 108(7): 1087–1111.
Murphy, B., Forest, F., Barraclough, T., Rosindell, J., Bellot, S., Cowan, R., … Cheek, M. (2020). A phylogenomic analysis of Nepenthes (Nepenthaceae). Molecular Phylogenetics and Evolution 144, 106668.
Ogutcen, E., Christe, C., Nishii, K., Salamin, N., Möller, M., Perret, M. (2021). Phylogenomics of Gesneriaceae using targeted capture of nuclear genes. Molecular Phylogenetics and Evolution 157, 107068.
Ottenlips, M.V., Mansfield, D.H., Buerki, S., Feist, M.A.E., Downie, S.R., Dodsworth, … Smith, J.F. (2021). Resolving species boundaries in a recent radiation with the Angiosperms353 probe set: the Lomatium packardiae/L. anomalum clade of the L. triternatum (Apiaceae) complex. American Journal of Botany 108, 1217–1233.
Pérez-Escobar, O.A., Dodsworth, S., Bogarín, D., Bellot, S., Balbuena, J.A., Schley, R.J., … Baker, W.J. (2021). Hundreds of nuclear and plastid loci yield novel insights into orchid relationships. American Journal of Botany 108, 1166–1180.
Pillon, Y., Hopkins, H.C.F., Maurin, O., Epitawalage, N., Bradford, J., Rogers, Z.S., … Forest, F. (2021). Phylogenomics and biogeography of Cunoniaceae (Oxalidales) with complete generic sampling and taxonomic realignments. American Journal of Botany 108, 1181–1200.
Shah, T., Schneider, J.V., Zizka, G., Maurin, O., Baker, W., Forest, F., … Larridon, I., (2021). Joining forces in Ochnaceae phylogenomics: a tale of two targeted sequencing probe kits. American Journal of Botany 108, 1201–1216.
Shee, Z. Q., D. G. Frodin, R. Cámara-Leret, L. Pokorny. (2020). Reconstructing the Complex Evolutionary History of the Papuasian Schefflera Radiation Through Herbariomics. Frontiers in Plant Science 11: 258.
Siniscalchi, C.M., Hidalgo, O., Palazzesi, L., Pellicer, J., Pokorny, L., Maurin, O., … Mandel, J.R. (2021). Lineage-specific vs. universal: A comparison of the Compositae1061 and Angiosperms353 enrichment panels in the sunflower family. Applications in Plant Sciences 9.
Starr, J.R., Jiménez-Mejías, P., Zuntini, A.R., Léveillé-Bourret, É., Semmouri, I., Muasya, M., … Larridon, I. (2021). Targeted sequencing supports morphology and embryo features in resolving the classification of Cyperaceae tribe Fuireneae s.l. Journal of Systematics and Evolution 59, 809–832.
Thomas, A.E., Igea, J., Meudt, H.M., Albach, D.C., Lee, W.G., Tanentzap, A.J. (2021). Using target sequence capture to improve the phylogenetic resolution of a rapid radiation in New Zealand Veronica. American Journal of Botany 108, 1289–1306.
Thomas, S.K., Liu, X., Du, Z.-Y., Dong, Y., Cummings, A., Pokorny, L., … Leebens-Mack, J.H. (2021). Comprehending Cornales: phylogenetic reconstruction of the order using the Angiosperms353 probe set. American Journal of Botany 108, 1112–1121.
Ufimov, R., Zeisek, V., Píšová, S., Baker, W.J., Fér, T., Loo, M., … Schmickl, R. (2021). Relative performance of customized and universal probe sets in target enrichment: A case study in subtribe Malinae. Applications in Plant Sciences 9, e11442.
Van Andel, T., Veltman, M. A., Bertin, A., Maat, H., Polime, T., Hille Ris Lambers, D., … Manzanilla, V.. (2019). Hidden Rice Diversity in the Guianas. Frontiers in Plant Science 10: 1161.
Wenzell, K.E., McDonnell, A.J., Wickett, N.J., Fant, J.B., Skogen, K.A. (2021). Incomplete reproductive isolation and low genetic differentiation despite floral divergence across varying geographic scales in Castilleja. American Journal of Botany 108, 1270–1288.
Zuntini, A.R., Frankel, L.P., Pokorny, L., Forest, F., Baker, W.J. (2021). A comprehensive phylogenomic study of the monocot order Commelinales, with a new classification of Commelinaceae. American Journal of Botany 108, 1066–1086.








